Tethered to science
For a long while I haven’t posted anything, being too tired and work got in the way. Then after combing back from a conference and a break late February, full of ideas but no energy to work them out, I turned to my GP. There I was told I was overworked, warned that if I did not slow down I will end up with a burn out. So I tried to rest and slow down, while still having that nudge of guild when leaving work early. Then the corona crisis hit. Forced to work at home made me feel less guilty for the times I stopped earlier, because hey we where in a crisis, so its ok not to be able to concentrate. Needless to say it did wonders for my recovery.
Working on a review article that was already in the pipeline, I recovered slowly. Not only am I again able to work a full day without loosing my concentration. I also found back what I lost long ago. How much I enjoyed just reading articles, being able to follow up on what I read, while discovering how it all worked. Then finding a way to describe this so others would see the connections between the different studies as well.
In short it showed me that biology is amazing and has find some inventive solutions to its problems.
One of these are tethering proteins. These are proteins that are connected to membranes of different organelles, say the endoplasmic reticulum (ER) and the plasma membrane, to keep them close to each other. Tethering proteins make use of two different domains. On one end of the protein they have a transmembrane domain. This domain insert itself through the membrane to anchor the protein to it. On the other end they have a couple of domains that can bind membranes.
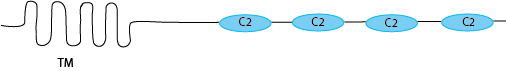

There are lots of different types of domains that can do that. But there is one in particular that is quite interesting in this context, the C2 domain. The C2 domain binds to membranes in a calcium dependent manner. But the concentration of calcium required depends on the exact sequence, so there are C2 domains that need just a little bit of calcium and they will already bind membranes. Then there are others that need quite a high calcium concentration for them even have a chance to bind the membrane. Tethering proteins make use of these C2 domain characteristics. They have a low calcium requirement C2 domain at the extreme end of the protein, and then another, one, two or three C2 domains, each requiring a bit more calcium before they are able to bind the membrane, placed more towards the transmembrane domain.

In this way, by using tethering proteins, the cell can hold both, say the ER and the plasma membrane, just having a little bit of calcium present.

But when the ER and the plasma membrane needs to be really close, say for some signalling function, then the cell can increase the calcium concentration, and the tethering protein will just reel in the ER close to the plasma membrane.
That image of the ER tethered via a leach to the plasma membrane, so it can be brought in close when needed. Is just one of the many I got over the past weeks, reading for my review. I will share some more over the coming weeks, illustrating how science reeled me in once again.